Hong Kong's marine waters are a true treasure trove of life. Despite the limited size of our sea area and heavy urban pressures, these waters support an astonishing biodiversity—around 6,000 known marine species, including more than 1,100 kinds of fish from 148 families, making up roughly 30% of the fish diversity found across the entire South China Sea. Our subtropical estuaries, shaped by the mixing of Pearl River freshwater and oceanic currents, create diverse habitats that serve as vital feeding and nursery grounds for numerous fish, marine mammals, and other creatures.
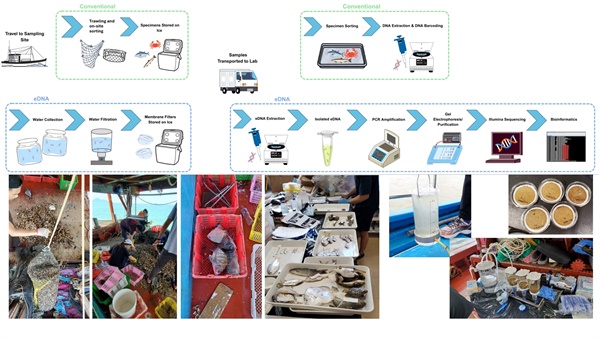
Historically, scientists relied on conventional fishing methods such as bottom trawling—dragging heavy nets across the seabed—to survey the underwater world. While effective in some respects, this method has serious drawbacks: it damaged sensitive habitats, is expensive and labour-intensive, and often overlooks species that reside higher in the water column, hide in complex environments, or are simply rare and elusive. Following the full bottom trawl ban took effect in late 2012, the need for gentler, more comprehensive monitoring methods has grown imperative for assessing recovery and guiding conservation efforts.
Environmental DNA (eDNA) metabarcoding presents a game-changing solution. Rather than catching animals, the eDNA approach captures tiny traces of genetic material that organisms naturally shed into seawater. By filtering water samples and analysing the DNA with advanced genetic techniques, we can identify species quickly, non-destructively, and with remarkable detail—no nets, no harm, and no bias against shy or mid-water species.
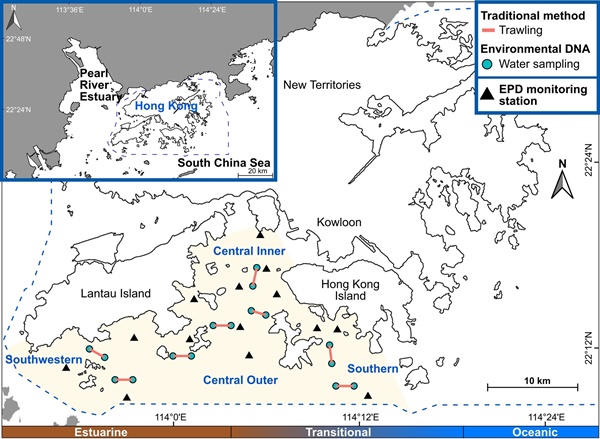
In our study, the first of its kind in Hong Kong's urbanised estuarine waters, we directly compared eDNA metabarcoding with traditional bottom trawling. In April and May 2022, we carried out 16 trawl hauls and collected 32 water samples across the southern waters of Hong Kong. Using multiple targeted genetic markers and high-throughput sequencing, we uncovered a much richer picture of the biodiversity present.
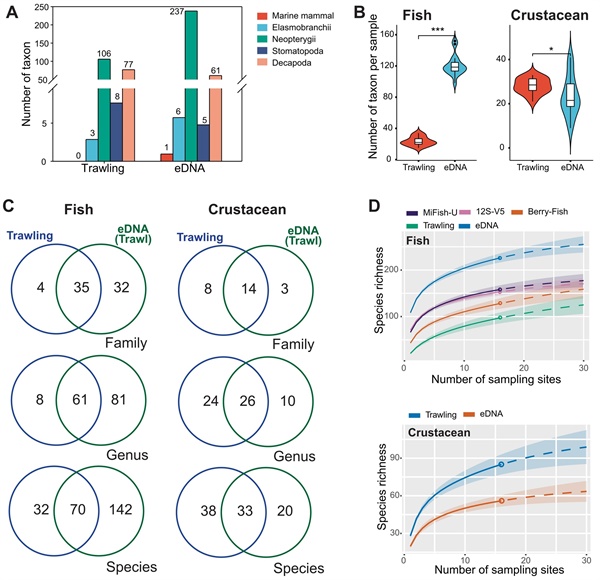
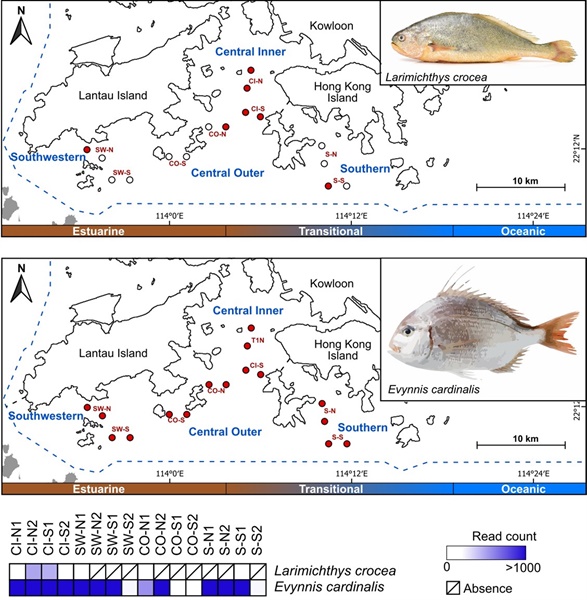
The results were compelling. Trawling detected 106 fish species and 3 rays, while eDNA identified 237 fish species and 6 rays—more than double the count, including 142 fish species that were completely missed by the nets. The eDNA approach also detected far more threatened vertebrates (16 compared to just 4 from trawling), such as the Critically Endangered Large Yellow Croaker (observed at over half the sites), the Endangered Evynnis cardinalis (found at every site), as well as the Vulnerable Indo-Pacific Finless Porpoise. The eDNA approach detected not only more fish species, but also clearer spatial patterns: distinct fish communities in different areas were closely linked to water quality factors like turbidity, oxygen levels, and nutrient concentrations.
Although trawling caught slightly more crustaceans, the eDNA approach still identified many key species, and improvements to genetic databases promise even better results in the future. These findings highlight eDNA metabarcoding's potential as a faster, less destructive, and more comprehensive approach for understanding Hong Kong's marine life. By revealing hidden diversity and tracking ecological changes without harming ecosystems, this innovative approach supports effective conservation, helps monitor recovery after the trawl ban, protects endangered species, and informs sustainable management of our valuable marine heritage.
Full paper: https://doi.org/10.1002/edn3.70031
| Principal Investigators | Prof. Jian-Wen QIU, Dr. Jack Chi-Ho IP |
|---|---|
| Affiliation | Hong Kong Baptist University |
| Co-investigators | Prof. Kenneth Mei Yee LEUNG, Dr. Leo Lai Chan, Mr. Vincent Chi-Sing Lai |
| Period | 2021–2024 |
| Funding Source | Lantau Conservation Fund |
Information Source: Prof. Jian-Wen Qiu, Dr. Jack Chi-Ho Ip